|
Ligand Docking
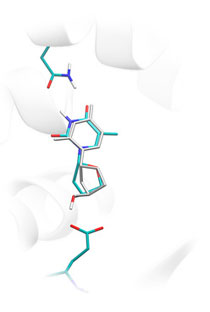
Lead Finder predicts 3D structure of non-covalent and covalently bound protein-ligand complexes by docking a flexible ligand to a static protein structure. The remarkable efficiency of Lead Finder is based on a unique conformational search algorithm and high accuracy scoring function to evaluate the free energy of protein-ligand binding (see the Science section). Accuracy of protein-ligand docking has been validated on a set of 407 protein-ligand complexes that appears to be the most extensive benchmarking study of such kind. This test set comprises test sets of such docking programs as
FlexX,
Glide SP and Glide XP,
Gold,
LigandFit,
MolDock,
Surflex
that allows fair and straightforward comparison of Lead Finder to the original predictions by the competitive programs. As can be seen from our benchmarking study, Lead Finder has outperformed all competitive programs on their native test sets, with docking success rates ranging from 80.0% on Glide XP and FlexX test sets to 96.0% on Surflex and MolDock test sets. Lead Finder has intuitive program interface that is easy to configure and use. While many configuration options are available to end users to fine-tune docking calculations, two calculation regimes, docking and screening, have been created to provide the optimum balance between the speed of calculations and the accuracy of predictions. Docking regime is slower but a bit more accurate. Screening regime is fine-tuned to achieve fastest performance at the cost of slight decrease in docking accuracy (see the Computing Speed section for more details). The speed of docking calculations has been extensively benchmarked both on the test set of 407 experimentally characterized protein-ligand complexes and in real-life virtual screening experiments of large commercial libraries of compounds against various drug targets. In our trials, docking of a single compound has taken 30-60 seconds on average in the more accurate docking regime, and 10-20 seconds in the faster virtual screening regime (see the Computing Speed section for more details).  |